Translational changes induced by acute sleep deprivation uncovered by TRAP- Seq | Molecular Brain | Full Text

TRAP-SEQ – Translating Ribosome Affinity Purification (TRAP) Followed by RNA Sequencing Technology | RNA-Seq Blog

Translational changes induced by acute sleep deprivation uncovered by TRAP- Seq | Molecular Brain | Full Text

Rapid Changes in the Translatome during the Conversion of Growth Cones to Synaptic Terminals - ScienceDirect

Evaluation of TRAP-seq by molecular profiling of the Emx1-lineage in... | Download Scientific Diagram

Translational changes induced by acute sleep deprivation uncovered by TRAP- Seq | Molecular Brain | Full Text
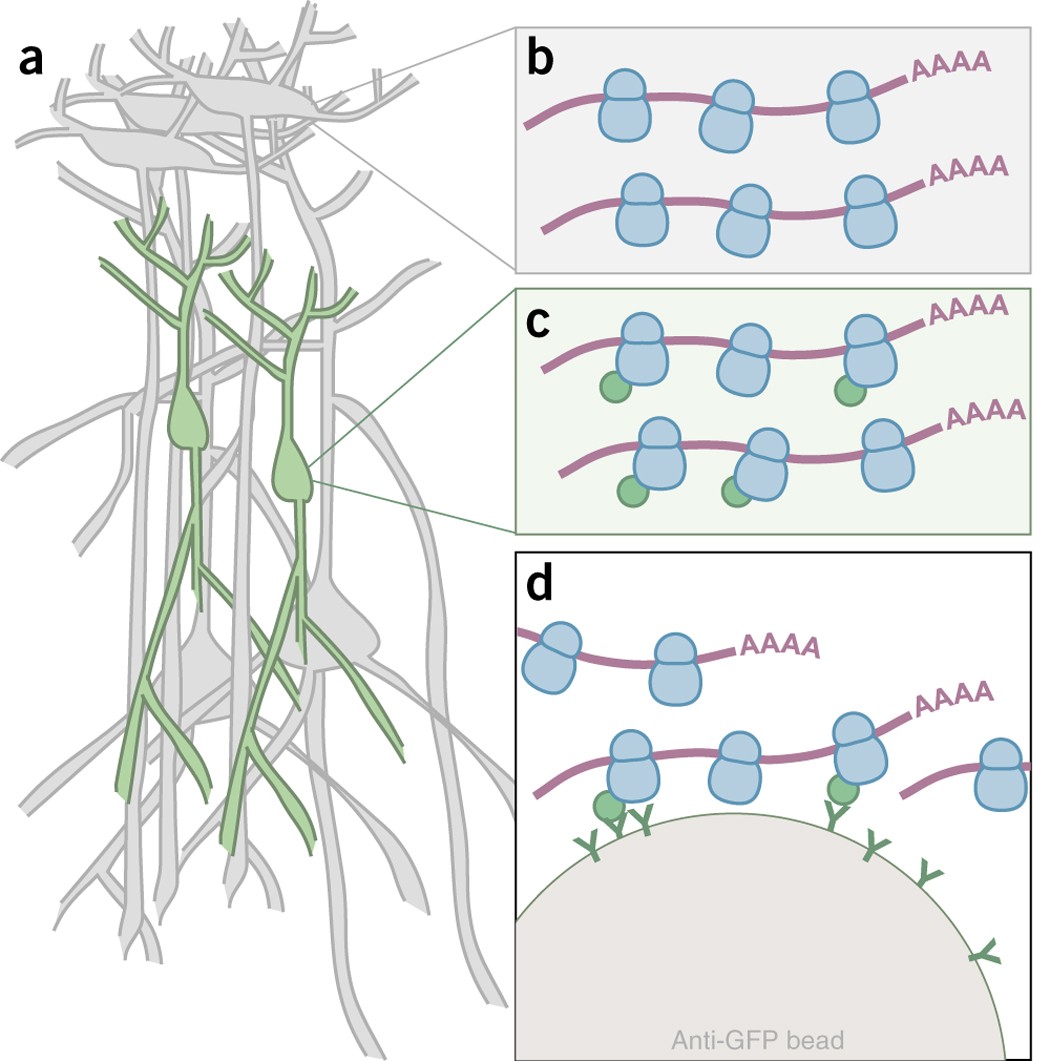
Cell type–specific mRNA purification by translating ribosome affinity purification (TRAP) | Nature Protocols
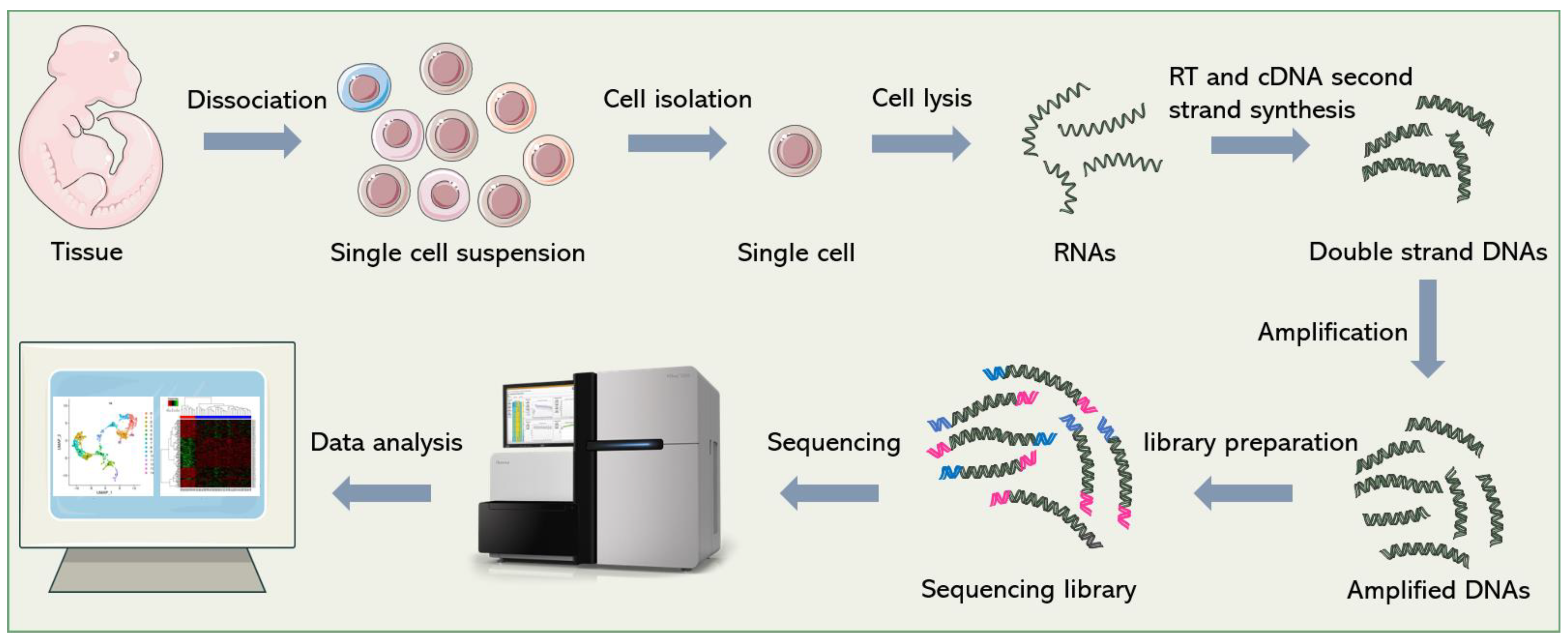
Biosensors | Free Full-Text | Microfluidics Facilitates the Development of Single-Cell RNA Sequencing
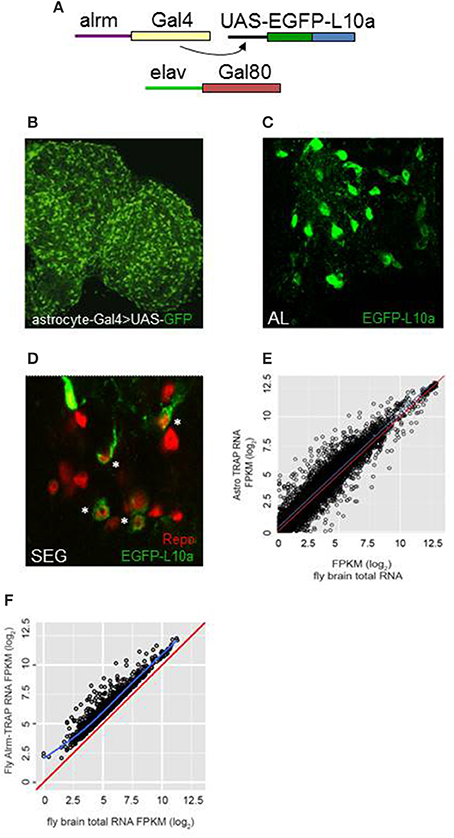
Frontiers | TRAP-seq Profiling and RNAi-Based Genetic Screens Identify Conserved Glial Genes Required for Adult Drosophila Behavior

TRAP-SEQ of Eukaryotic Translatomes Applied to the Detection of Polysome-Associated Long Noncoding RNAs | SpringerLink

Translating Ribosome Affinity Purification (TRAP) Followed by RNA Sequencing Technology (TRAP-SEQ) for Quantitative Assessment of Plant Translatomes | SpringerLink
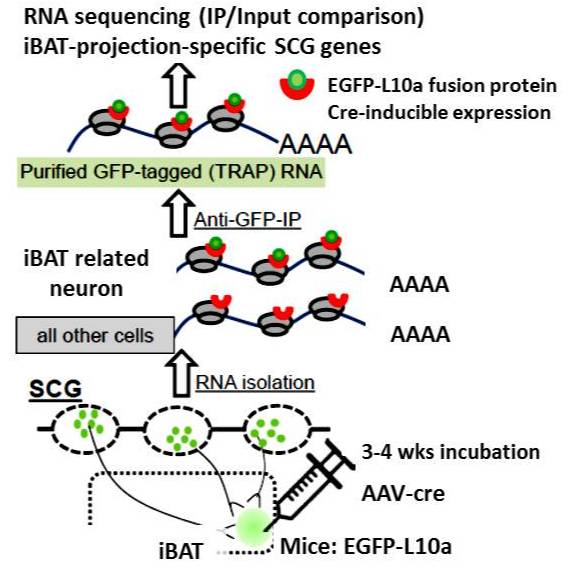
TRAP-SEQ (Translating Ribosome Affinity Purification followed by RNA sequencing) of interscapular brown adipose tissue (iBAT)- related ganglia from 7-day cold and warm treated mice - Blackfynn Discover